# SciELO XML to PubMed XML
Program to export XML to PubMed, according to [http://www.ncbi.nlm.nih.gov/books/NBK3828/](http://www.ncbi.nlm.nih.gov/books/NBK3828/), using SciELO XML (SciELO Publishing Schema).
## How to execute
Double clicking on c:\\scielo\\bin\\xml\\xml\_pubmed.py
> 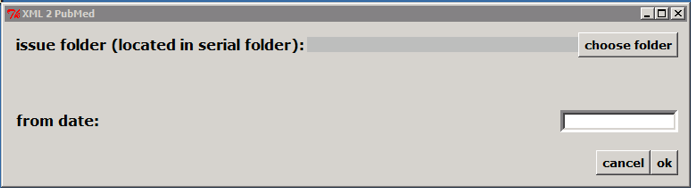
Select the issue folder
> 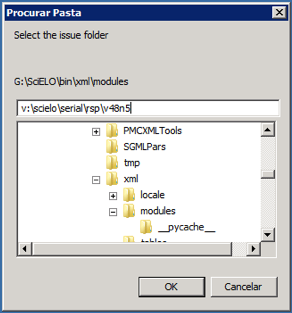
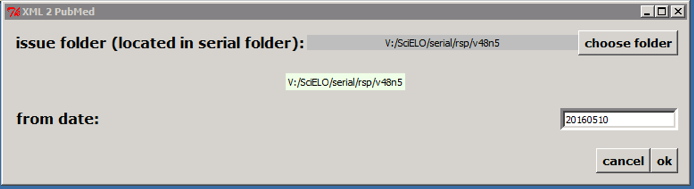
Only if issue is published on batches, such aop or rolling pass, you should inform **from date** to generate XML for the article published from this date to the current date.
Then click on OK button.
According to the example, the program will create the file: v:\\scielo\\serial\\rsp\\v48n5\\PubMed\\rsp-v48n5-20160510-20160523.xml, containing articles which have epub date between 20160510 and the current date.
> 
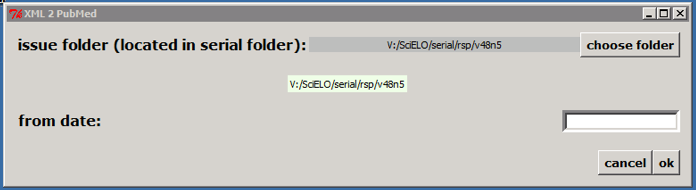
If it is not an issue published on batches, click on OK button. According to the example, the program will create the file: v:\\scielo\\serial\\rsp\\v48n5\\PubMed\\rsp-v48n5.xml.
> 
Or execute it on a terminal:
> 
Optionally informing the **from date**
> 